Jump to Research, Methods, Tools.
Research
RNA transcription is the first step in the expression of genetic information and is highly regulated during cell differentiation and organism development. Understanding how transcription is regulated in health and disease will ultimately lead to new diagnostic tools and therapies for genetic disorders.
We put a major emphasis on understanding the dynamics of transcription. Our unique approach allows us to define new regulatory elements by characterizing biophysical properties of DNA and RNA. We apply this methodology to poorly characterized transcription systems such as SARS-CoV-2 RNA polymerase and RNA polymerase III in humans. Here are some themes and techniques that we currently work on:
RNA-dependent RNA polymerase of SARS-CoV-2 coronavirus
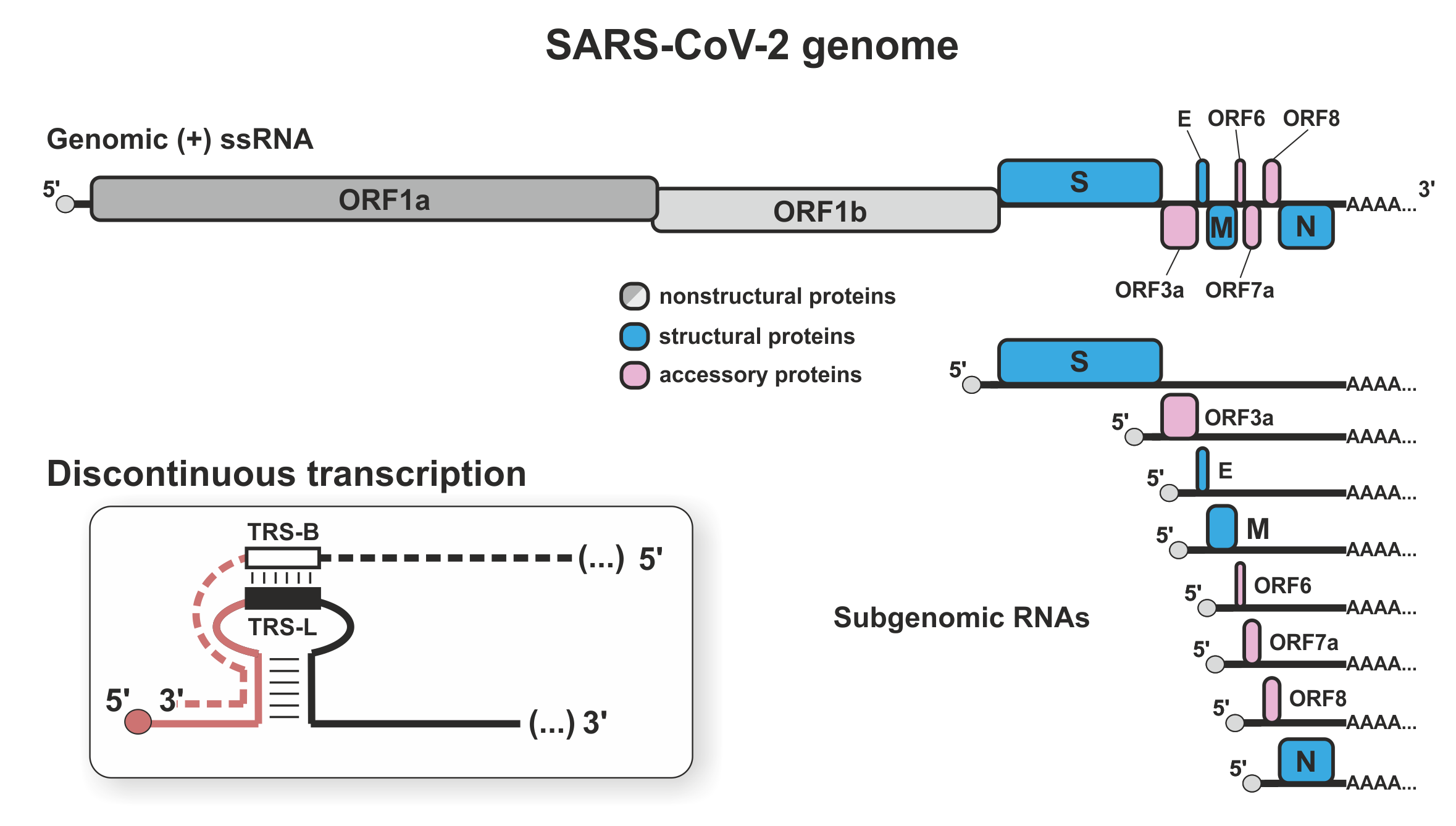
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a positive strand RNA virus, in which the infecting RNA serves both as mRNA and genome. An RNA-dependent RNA polymerase (RdRP) both transcribes the genes and replicates the genome, making RdRP a major target in COVID19 therapy. Nucleotide analogues, including the drugs remdesivir and sofosbuvir, inhibit RNA synthesis by RdRP, but treatments could be enhanced by better understanding the key features of RdRP function.
Within SARS-CoV-2 infected cells, abundant structural proteins are synthesised from subgenomic RNA fragments (sgRNAs) generated by discontinuous transcription. Elongating RdRP can ‘jump’ across large regions of the genome before resuming transcription. The abundance of the sgRNAs controls the stoichiometry between viral proteins. The jump mechanism is unclear, but transcription elongation rates, backtracking, and premature dissociation seem likely factors. It is also unclear how RdRP switches between discontinuous transcription and continuous replication. We use our experience in determining transcription mechanisms in eukaryotic cells to resolve these key questions.
To define key elements in SARS-CoV-2 transcription regulation, we are applying a combination of in vivo biology, biochemistry, bioinformatics and mathematical modelling. Identification of sequences and mechanisms important for discontinuous transcription will help clarify inhibitory mechanisms triggered by antivirals, allowing directed improvements and the identification of potentially synergistic therapeutic targets.
Molecular mechanisms linking RNA polymerase III mutations to neurodegenerative diseases
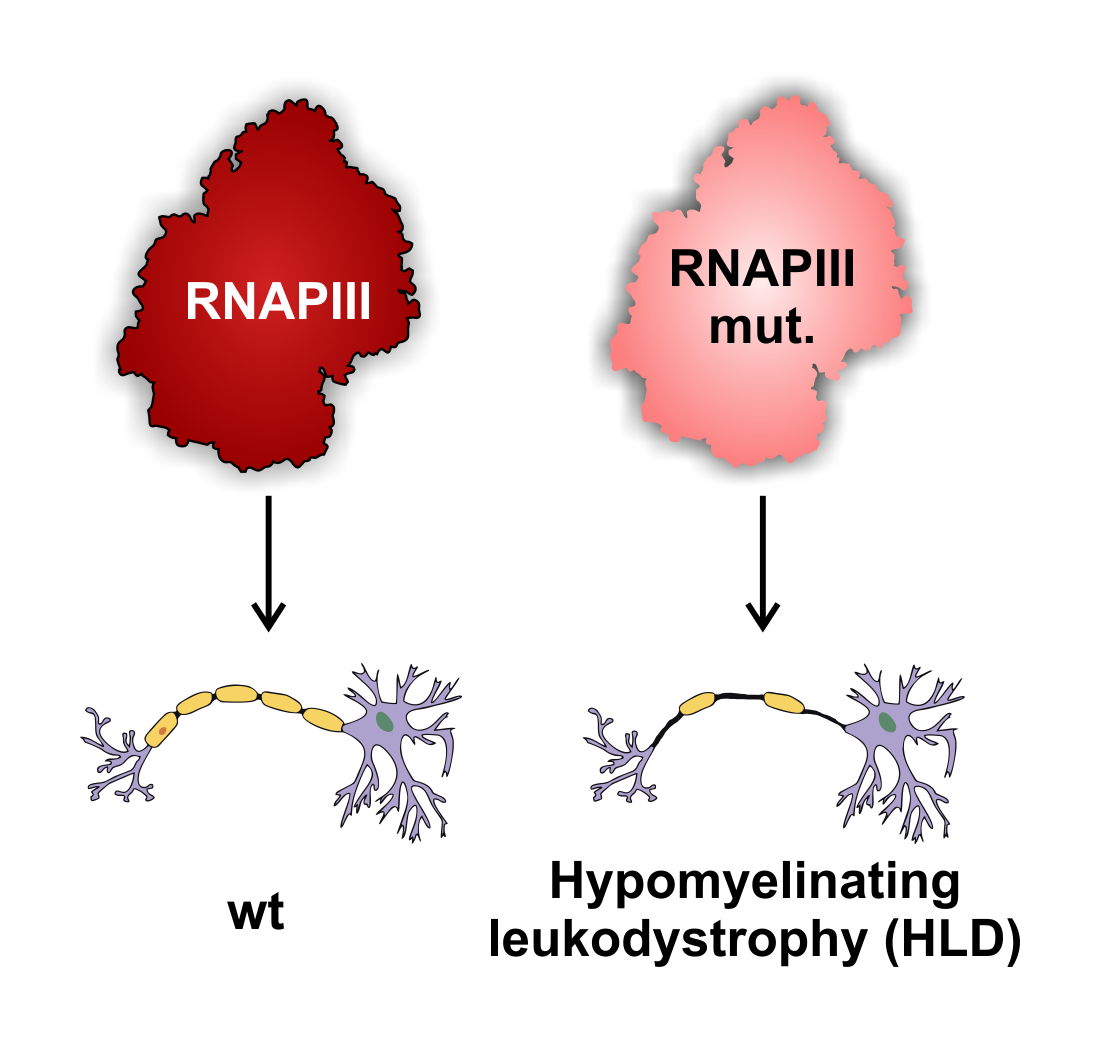
The human genome is transcribed by three RNA polymerases (RNAPs): RNAPI, RNAPII and RNAPIII. Protein coding genes and many non-coding genes are transcribed by RNAPII, whereas ribosomal RNA (rRNA) and transfer RNAs (tRNAs) are transcribed by RNAPI and III, respectively. RNAPIII also transcribes several small non-coding RNAs that are crucial for cell viability. Most of the RNAPIII mutations are predicted to be lethal, whereas others cause tissue-specific phenotypes.
We are trying to identify the role of RNAPIII in the molecular mechanisms underlying a group of neurodegenerative diseases associated with loss of myelination, known as Hypomyelinating leukodystrophies (HLD). A common feature of all leukodystrophies is degeneration of white matter in the brain caused by defects in the myelin sheath around axons. Myelin is formed by oligodendrocyte cells during differentiation of oligodendrocyte progenitor cells.
To understand the molecular phenotype of these disorders, we are mapping actively transcribing RNAPIII and measuring the pool of tRNA available for translation. Additionally, we are investigating the effect of RNAPIII transcription on the expression of surrounding protein-coding genes.
This research will provide insights into the role of RNAPIII mutations in HLD. We aim to define transcripts that are directly affected by compromised RNAPIII activity or are particularly sensitive during oligodendrocyte differentiation. We envisage that our predictions can be tested in disease-relevant models to shed new light on the molecular mechanism of RNAPIII-related HLD.
Methods
Biological models
Culturing and genetic manipulations including CRISPR-based approches.
- Cell lines: K562, A549, HeLa, Caco-2, Calu-1, LR7
- Mouse embrionic stem cells (mESC)
- Budding yeast (Saccharomyces cerevisiae)

High-throughput methods
- CRAC (UV cross-linking and analysis of cDNA), method that provides high-resolution maps of RNA-protein interactions. We use CRAC to investigate RNA binding proteins and the dynamics of RNA transcription
- ChIP-nexus to investigate DNA-protein interactions, a high-resolution modification of the chromatin immunoprecipitation (ChIP) method.
- RNA-seq
- tRNA-seq
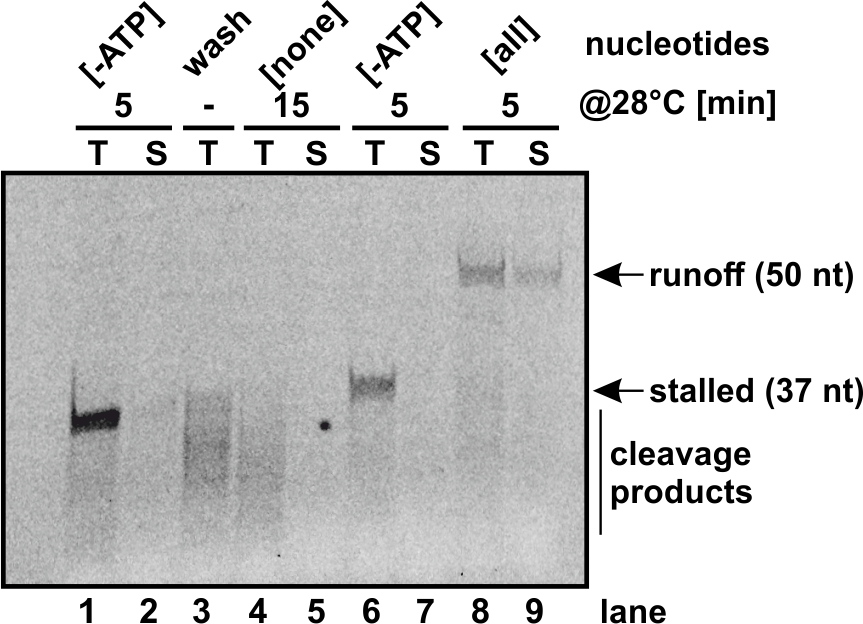
In vitro biochemistry
- Purification of large complexes for in vitro assays: RNA polymerases, pre-ribosomes etc
- RNA polymerase I and II in vitro assays: elongation and backtracking
- SARS-CoV-2 coronavirus RNA-dependent RNA polymerase elongation assay
- Cleavage assay of PIN-domain endonucleases (i.e. Nob1)
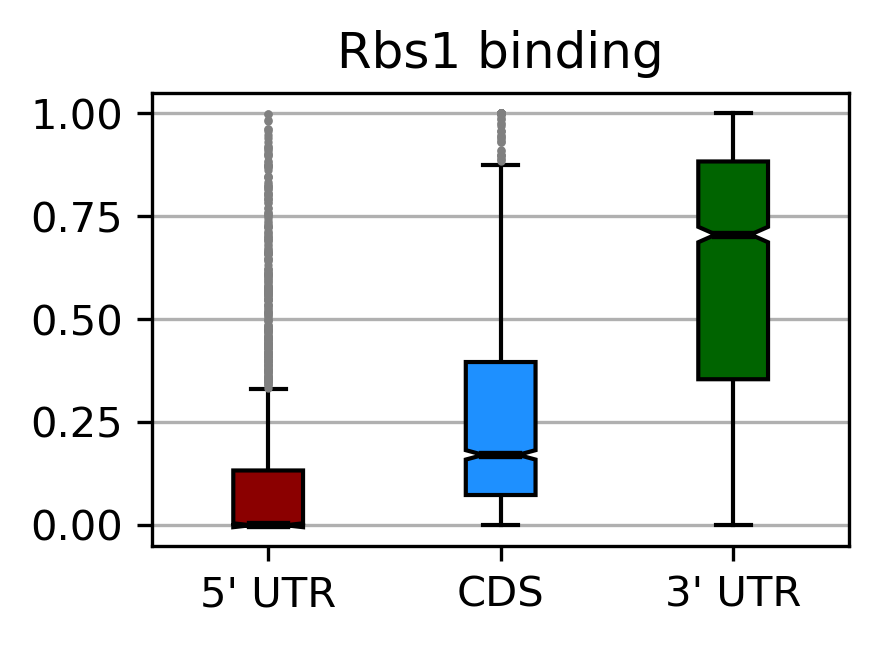
Data analysis and mathematical modelling:
- RNA-seq, tRNA-seq, CRAC/CLIP data analysis.
- Analysis of RNA polymerase dynamics
- Motif analysis
- Data science approach to cross-compare various types of datasets
- Mathematical modelling of dynamic processes with a major focus on transcription of RNA
- High-scale parametrization of DNA and RNA sequences to thermodynamic features (folding energy, melting energy, shape features etc.)
Tools
Python with Jupyter Notebook is our default working environment. We also use:
We also develope in-house tools, some of them are already publicly availible:
- trxtools tools. Package of python tools facilitating bioinformatic analysis of RNA and nascent RNA sequencing data. Check documentation for details.
- Mathematical model of RNAP. MATLAB code simulating kinetics of RNA polymerase I published in Turowski et al 2020 Molecular Cell paper.